| 1 | c3h2zA_
|
|
 |
100.0 |
97 |
PDB header:oxidoreductase
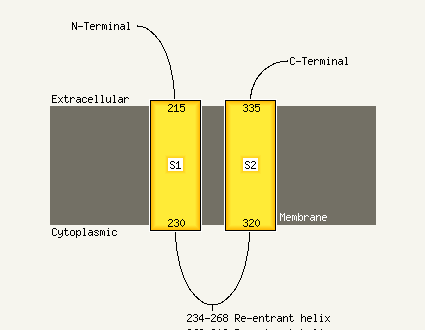
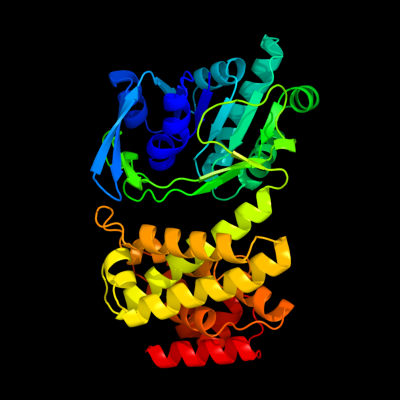
Chain: A: PDB Molecule:mannitol-1-phosphate 5-dehydrogenase;
PDBTitle: the crystal structure of mannitol-1-phosphate dehydrogenase from2 shigella flexneri
|
| 2 | c1m2wA_
|
|
 |
100.0 |
20 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:mannitol dehydrogenase;
PDBTitle: pseudomonas fluorescens mannitol 2-dehydrogenase ternary complex with2 nad and d-mannitol
|
| 3 | d1lj8a4
|
|
 |
100.0 |
19 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 4 | d1lj8a3
|
|
 |
100.0 |
24 |
Fold:6-phosphogluconate dehydrogenase C-terminal domain-like
Superfamily:6-phosphogluconate dehydrogenase C-terminal domain-like
Family:Mannitol 2-dehydrogenase |
| 5 | c3hn2A_
|
|
 |
98.1 |
19 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:2-dehydropantoate 2-reductase;
PDBTitle: crystal structure of 2-dehydropantoate 2-reductase from geobacter2 metallireducens gs-15
|
| 6 | c2ew2B_
|
|
 |
98.0 |
15 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:2-dehydropantoate 2-reductase, putative;
PDBTitle: crystal structure of the putative 2-dehydropantoate 2-reductase from2 enterococcus faecalis
|
| 7 | c3c7cB_
|
|
 |
97.9 |
16 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:octopine dehydrogenase;
PDBTitle: a structural basis for substrate and stereo selectivity in2 octopine dehydrogenase (odh-nadh-l-arginine)
|
| 8 | c3ghyA_
|
|
 |
97.6 |
19 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:ketopantoate reductase protein;
PDBTitle: crystal structure of a putative ketopantoate reductase from ralstonia2 solanacearum molk2
|
| 9 | c3egoB_
|
|
 |
97.5 |
16 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:probable 2-dehydropantoate 2-reductase;
PDBTitle: crystal structure of probable 2-dehydropantoate 2-reductase2 pane from bacillus subtilis
|
| 10 | c3hwrA_
|
|
 |
97.3 |
16 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:2-dehydropantoate 2-reductase;
PDBTitle: crystal structure of pane/apba family ketopantoate reductase2 (yp_299159.1) from ralstonia eutropha jmp134 at 2.15 a resolution
|
| 11 | c3k96B_
|
|
 |
97.1 |
13 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:glycerol-3-phosphate dehydrogenase [nad(p)+];
PDBTitle: 2.1 angstrom resolution crystal structure of glycerol-3-phosphate2 dehydrogenase (gpsa) from coxiella burnetii
|
| 12 | c1bg6A_
|
|
 |
97.1 |
17 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:n-(1-d-carboxylethyl)-l-norvaline dehydrogenase;
PDBTitle: crystal structure of the n-(1-d-carboxylethyl)-l-norvaline2 dehydrogenase from arthrobacter sp. strain 1c
|
| 13 | c2axqA_
|
|
 |
97.1 |
14 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:saccharopine dehydrogenase;
PDBTitle: apo histidine-tagged saccharopine dehydrogenase (l-glu2 forming) from saccharomyces cerevisiae
|
| 14 | d1mv8a2
|
|
 |
97.1 |
14 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 15 | c2qytA_
|
|
 |
97.0 |
19 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:2-dehydropantoate 2-reductase;
PDBTitle: crystal structure of 2-dehydropantoate 2-reductase from porphyromonas2 gingivalis w83
|
| 16 | c3g17H_
|
|
 |
96.8 |
15 |
PDB header:structural genomics, unknown function
Chain: H: PDB Molecule:similar to 2-dehydropantoate 2-reductase;
PDBTitle: structure of putative 2-dehydropantoate 2-reductase from2 staphylococcus aureus
|
| 17 | d1n1ea2
|
|
 |
96.8 |
13 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 18 | d1txga2
|
|
 |
96.6 |
12 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 19 | c3qhaB_
|
|
 |
96.6 |
17 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:putative oxidoreductase;
PDBTitle: crystal structure of a putative oxidoreductase from mycobacterium2 avium 104
|
| 20 | c1txgA_
|
|
 |
96.6 |
14 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glycerol-3-phosphate dehydrogenase [nad(p)+];
PDBTitle: structure of glycerol-3-phosphate dehydrogenase from archaeoglobus2 fulgidus
|
| 21 | c1m67A_ |
|
not modelled |
96.5 |
13 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glycerol-3-phosphate dehydrogenase;
PDBTitle: crystal structure of leishmania mexicana gpdh complexed with inhibitor2 2-bromo-6-hydroxy-purine
|
| 22 | c2ofpB_ |
|
not modelled |
96.4 |
16 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:ketopantoate reductase;
PDBTitle: crystal structure of escherichia coli ketopantoate2 reductase in a ternary complex with nadp+ and pantoate
|
| 23 | c3ic5A_ |
|
not modelled |
96.3 |
23 |
PDB header:structural genomics, unknown function
Chain: A: PDB Molecule:putative saccharopine dehydrogenase;
PDBTitle: n-terminal domain of putative saccharopine dehydrogenase from ruegeria2 pomeroyi.
|
| 24 | d1bg6a2 |
|
not modelled |
96.1 |
18 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 25 | c3i83B_ |
|
not modelled |
96.0 |
15 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:2-dehydropantoate 2-reductase;
PDBTitle: crystal structure of 2-dehydropantoate 2-reductase from methylococcus2 capsulatus
|
| 26 | c2f1kD_ |
|
not modelled |
95.8 |
16 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:prephenate dehydrogenase;
PDBTitle: crystal structure of synechocystis arogenate dehydrogenase
|
| 27 | c3l4bG_ |
|
not modelled |
95.5 |
5 |
PDB header:transport protein
Chain: G: PDB Molecule:trka k+ channel protien tm1088b;
PDBTitle: crystal structure of an octomeric two-subunit trka k+ channel ring2 gating assembly, tm1088a:tm1088b, from thermotoga maritima
|
| 28 | c2z2vA_ |
|
not modelled |
95.4 |
31 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:hypothetical protein ph1688;
PDBTitle: crystal structure of l-lysine dehydrogenase from2 hyperthermophilic archaeon pyrococcus horikoshii
|
| 29 | c2gf2B_ |
|
not modelled |
95.4 |
25 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:3-hydroxyisobutyrate dehydrogenase;
PDBTitle: crystal structure of human hydroxyisobutyrate dehydrogenase
|
| 30 | d2f1ka2 |
|
not modelled |
95.3 |
26 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 31 | c3d4oA_ |
|
not modelled |
95.3 |
14 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:dipicolinate synthase subunit a;
PDBTitle: crystal structure of dipicolinate synthase subunit a (np_243269.1)2 from bacillus halodurans at 2.10 a resolution
|
| 32 | d1jaya_ |
|
not modelled |
95.1 |
18 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 33 | c3l6dB_ |
|
not modelled |
95.1 |
17 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:putative oxidoreductase;
PDBTitle: crystal structure of putative oxidoreductase from pseudomonas putida2 kt2440
|
| 34 | c1e5lA_ |
|
not modelled |
94.7 |
21 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:saccharopine reductase;
PDBTitle: apo saccharopine reductase from magnaporthe grisea
|
| 35 | c2ep9A_ |
|
not modelled |
94.7 |
12 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:l-gulonate 3-dehydrogenase;
PDBTitle: crystal structure of the rabbit l-gulonate 3-dehydrogenase2 (nadh form)
|
| 36 | c3dojA_ |
|
not modelled |
94.6 |
23 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:dehydrogenase-like protein;
PDBTitle: structure of glyoxylate reductase 1 from arabidopsis2 (atglyr1)
|
| 37 | d1ks9a2 |
|
not modelled |
94.5 |
18 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 38 | d1lssa_ |
|
not modelled |
94.4 |
20 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Potassium channel NAD-binding domain |
| 39 | d2ahra2 |
|
not modelled |
94.3 |
22 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 40 | c3ckyA_ |
|
not modelled |
94.1 |
23 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:2-hydroxymethyl glutarate dehydrogenase;
PDBTitle: structural and kinetic properties of a beta-hydroxyacid dehydrogenase2 involved in nicotinate fermentation
|
| 41 | c3eywA_ |
|
not modelled |
94.1 |
18 |
PDB header:transport protein
Chain: A: PDB Molecule:c-terminal domain of glutathione-regulated potassium-efflux
PDBTitle: crystal structure of the c-terminal domain of e. coli kefc in complex2 with keff
|
| 42 | c3c24A_ |
|
not modelled |
94.0 |
23 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:putative oxidoreductase;
PDBTitle: crystal structure of a putative oxidoreductase (yp_511008.1) from2 jannaschia sp. ccs1 at 1.62 a resolution
|
| 43 | c3g0oA_ |
|
not modelled |
93.9 |
28 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:3-hydroxyisobutyrate dehydrogenase;
PDBTitle: crystal structure of 3-hydroxyisobutyrate dehydrogenase2 (ygbj) from salmonella typhimurium
|
| 44 | d1dlja2 |
|
not modelled |
93.9 |
30 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 45 | c1z82A_ |
|
not modelled |
93.8 |
23 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:glycerol-3-phosphate dehydrogenase;
PDBTitle: crystal structure of glycerol-3-phosphate dehydrogenase (tm0378) from2 thermotoga maritima at 2.00 a resolution
|
| 46 | c2uyyD_ |
|
not modelled |
93.3 |
23 |
PDB header:cytokine
Chain: D: PDB Molecule:n-pac protein;
PDBTitle: structure of the cytokine-like nuclear factor n-pac
|
| 47 | c2vhyB_ |
|
not modelled |
93.3 |
24 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:alanine dehydrogenase;
PDBTitle: crystal structure of apo l-alanine dehydrogenase from2 mycobacterium tuberculosis
|
| 48 | c3gg2B_ |
|
not modelled |
92.9 |
18 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:sugar dehydrogenase, udp-glucose/gdp-mannose
PDBTitle: crystal structure of udp-glucose 6-dehydrogenase from2 porphyromonas gingivalis bound to product udp-glucuronate
|
| 49 | c2vrcD_ |
|
not modelled |
92.7 |
17 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:triphenylmethane reductase;
PDBTitle: crystal structure of the citrobacter sp. triphenylmethane2 reductase complexed with nadp(h)
|
| 50 | c1pjcA_ |
|
not modelled |
92.5 |
24 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:protein (l-alanine dehydrogenase);
PDBTitle: l-alanine dehydrogenase complexed with nad
|
| 51 | c1gpjA_ |
|
not modelled |
92.5 |
23 |
PDB header:reductase
Chain: A: PDB Molecule:glutamyl-trna reductase;
PDBTitle: glutamyl-trna reductase from methanopyrus kandleri
|
| 52 | c2y0dB_ |
|
not modelled |
92.3 |
33 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:udp-glucose dehydrogenase;
PDBTitle: bcec mutation y10k
|
| 53 | d2hmva1 |
|
not modelled |
92.3 |
20 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Potassium channel NAD-binding domain |
| 54 | d1l7da1 |
|
not modelled |
92.1 |
17 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
| 55 | c1vpdA_ |
|
not modelled |
92.0 |
26 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:tartronate semialdehyde reductase;
PDBTitle: x-ray crystal structure of tartronate semialdehyde reductase2 [salmonella typhimurium lt2]
|
| 56 | c3ktdC_ |
|
not modelled |
91.5 |
17 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:prephenate dehydrogenase;
PDBTitle: crystal structure of a putative prephenate dehydrogenase (cgl0226)2 from corynebacterium glutamicum atcc 13032 at 2.60 a resolution
|
| 57 | c3dttA_ |
|
not modelled |
91.5 |
20 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:nadp oxidoreductase;
PDBTitle: crystal structure of a putative f420 dependent nadp-reductase2 (arth_0613) from arthrobacter sp. fb24 at 1.70 a resolution
|
| 58 | d1pgja2 |
|
not modelled |
91.3 |
12 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 59 | c2ahrB_ |
|
not modelled |
91.1 |
23 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:putative pyrroline carboxylate reductase;
PDBTitle: crystal structures of 1-pyrroline-5-carboxylate reductase from human2 pathogen streptococcus pyogenes
|
| 60 | c3prjB_ |
|
not modelled |
91.0 |
21 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:udp-glucose 6-dehydrogenase;
PDBTitle: role of packing defects in the evolution of allostery and induced fit2 in human udp-glucose dehydrogenase.
|
| 61 | c3cumA_ |
|
not modelled |
90.9 |
30 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:probable 3-hydroxyisobutyrate dehydrogenase;
PDBTitle: crystal structure of a possible 3-hydroxyisobutyrate dehydrogenase2 from pseudomonas aeruginosa pao1
|
| 62 | c1mv8A_ |
|
not modelled |
90.7 |
27 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:gdp-mannose 6-dehydrogenase;
PDBTitle: 1.55 a crystal structure of a ternary complex of gdp-mannose2 dehydrogenase from psuedomonas aeruginosa
|
| 63 | c3plnA_ |
|
not modelled |
90.2 |
30 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:udp-glucose 6-dehydrogenase;
PDBTitle: crystal structure of klebsiella pneumoniae udp-glucose 6-dehydrogenase2 complexed with udp-glucose
|
| 64 | c3euwB_ |
|
not modelled |
90.2 |
26 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:myo-inositol dehydrogenase;
PDBTitle: crystal structure of a myo-inositol dehydrogenase from corynebacterium2 glutamicum atcc 13032
|
| 65 | d1obba1 |
|
not modelled |
90.1 |
23 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:LDH N-terminal domain-like |
| 66 | c3b1fA_ |
|
not modelled |
90.1 |
18 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:putative prephenate dehydrogenase;
PDBTitle: crystal structure of prephenate dehydrogenase from streptococcus2 mutans
|
| 67 | c1i36A_ |
|
not modelled |
89.9 |
18 |
PDB header:structural genomics, unknown function
Chain: A: PDB Molecule:conserved hypothetical protein mth1747;
PDBTitle: structure of conserved protein mth1747 of unknown function2 reveals structural similarity with 3-hydroxyacid3 dehydrogenases
|
| 68 | d1uxja1 |
|
not modelled |
89.8 |
15 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:LDH N-terminal domain-like |
| 69 | c3ezyB_ |
|
not modelled |
89.6 |
21 |
PDB header:structural genomics, unknown function
Chain: B: PDB Molecule:dehydrogenase;
PDBTitle: crystal structure of probable dehydrogenase tm_0414 from2 thermotoga maritima
|
| 70 | c2w2kB_ |
|
not modelled |
89.5 |
33 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:d-mandelate dehydrogenase;
PDBTitle: crystal structure of the apo forms of rhodotorula graminis2 d-mandelate dehydrogenase at 1.8a.
|
| 71 | c1ks9A_ |
|
not modelled |
88.9 |
15 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:2-dehydropantoate 2-reductase;
PDBTitle: ketopantoate reductase from escherichia coli
|
| 72 | c2rirA_ |
|
not modelled |
88.8 |
17 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:dipicolinate synthase, a chain;
PDBTitle: crystal structure of dipicolinate synthase, a chain, from bacillus2 subtilis
|
| 73 | d1i36a2 |
|
not modelled |
88.8 |
18 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 74 | c2izzE_ |
|
not modelled |
88.7 |
16 |
PDB header:oxidoreductase
Chain: E: PDB Molecule:pyrroline-5-carboxylate reductase 1;
PDBTitle: crystal structure of human pyrroline-5-carboxylate2 reductase
|
| 75 | d1pjca1 |
|
not modelled |
88.7 |
24 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Formate/glycerate dehydrogenases, NAD-domain |
| 76 | d1yqga2 |
|
not modelled |
88.4 |
15 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 77 | c2x4gA_ |
|
not modelled |
88.3 |
23 |
PDB header:isomerase
Chain: A: PDB Molecule:nucleoside-diphosphate-sugar epimerase;
PDBTitle: crystal structure of pa4631, a nucleoside-diphosphate-sugar2 epimerase from pseudomonas aeruginosa
|
| 78 | c2g5cD_ |
|
not modelled |
88.3 |
20 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:prephenate dehydrogenase;
PDBTitle: crystal structure of prephenate dehydrogenase from aquifex aeolicus
|
| 79 | c2vq3B_ |
|
not modelled |
88.1 |
19 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:metalloreductase steap3;
PDBTitle: crystal structure of the membrane proximal oxidoreductase2 domain of human steap3, the dominant ferric reductase of3 the erythroid transferrin cycle
|
| 80 | c3llvA_ |
|
not modelled |
88.0 |
21 |
PDB header:nad(p) binding protein
Chain: A: PDB Molecule:exopolyphosphatase-related protein;
PDBTitle: the crystal structure of the nad(p)-binding domain of an2 exopolyphosphatase-related protein from archaeoglobus fulgidus to3 1.7a
|
| 81 | c1dliA_ |
|
not modelled |
88.0 |
30 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:udp-glucose dehydrogenase;
PDBTitle: the first structure of udp-glucose dehydrogenase (udpgdh) reveals the2 catalytic residues necessary for the two-fold oxidation
|
| 82 | d1gpja2 |
|
not modelled |
88.0 |
22 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Aminoacid dehydrogenase-like, C-terminal domain |
| 83 | c2eezG_ |
|
not modelled |
87.9 |
25 |
PDB header:oxidoreductase
Chain: G: PDB Molecule:alanine dehydrogenase;
PDBTitle: crystal structure of alanine dehydrogenase from themus thermophilus
|
| 84 | c3iusB_ |
|
not modelled |
87.6 |
24 |
PDB header:structural genomics, unknown function
Chain: B: PDB Molecule:uncharacterized conserved protein;
PDBTitle: the structure of a functionally unknown conserved protein2 from silicibacter pomeroyi dss
|
| 85 | c2o3jC_ |
|
not modelled |
87.6 |
14 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:udp-glucose 6-dehydrogenase;
PDBTitle: structure of caenorhabditis elegans udp-glucose dehydrogenase
|
| 86 | c3pefA_ |
|
not modelled |
87.4 |
24 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:6-phosphogluconate dehydrogenase, nad-binding;
PDBTitle: crystal structure of gamma-hydroxybutyrate dehydrogenase from2 geobacter metallireducens in complex with nadp+
|
| 87 | c3icpA_ |
|
not modelled |
87.3 |
30 |
PDB header:isomerase
Chain: A: PDB Molecule:nad-dependent epimerase/dehydratase;
PDBTitle: crystal structure of udp-galactose 4-epimerase
|
| 88 | c1obbB_ |
|
not modelled |
87.1 |
23 |
PDB header:hydrolase
Chain: B: PDB Molecule:alpha-glucosidase;
PDBTitle: alpha-glucosidase a, agla, from thermotoga maritima in2 complex with maltose and nad+
|
| 89 | c3ceaA_ |
|
not modelled |
87.1 |
21 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:myo-inositol 2-dehydrogenase;
PDBTitle: crystal structure of myo-inositol 2-dehydrogenase (np_786804.1) from2 lactobacillus plantarum at 2.40 a resolution
|
| 90 | d1vl0a_ |
|
not modelled |
86.9 |
30 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Tyrosine-dependent oxidoreductases |
| 91 | c3h2sA_ |
|
not modelled |
86.3 |
13 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:putative nadh-flavin reductase;
PDBTitle: crystal structure of the q03b84 protein from lactobacillus2 casei. northeast structural genomics consortium target3 lcr19.
|
| 92 | c3qvoA_ |
|
not modelled |
86.3 |
16 |
PDB header:structural genomics, unknown function
Chain: A: PDB Molecule:nmra family protein;
PDBTitle: structure of a rossmann-fold nad(p)-binding family protein from2 shigella flexneri.
|
| 93 | c3gpiA_ |
|
not modelled |
86.3 |
27 |
PDB header:structural genomics, unknown function
Chain: A: PDB Molecule:nad-dependent epimerase/dehydratase;
PDBTitle: structure of putative nad-dependent epimerase/dehydratase2 from methylobacillus flagellatus
|
| 94 | c3dhnA_ |
|
not modelled |
85.8 |
27 |
PDB header:isomerase, lyase
Chain: A: PDB Molecule:nad-dependent epimerase/dehydratase;
PDBTitle: crystal structure of the putative epimerase q89z24_bactn2 from bacteroides thetaiotaomicron. northeast structural3 genomics consortium target btr310.
|
| 95 | c3d1lB_ |
|
not modelled |
85.6 |
16 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:putative nadp oxidoreductase bf3122;
PDBTitle: crystal structure of putative nadp oxidoreductase bf3122 from2 bacteroides fragilis
|
| 96 | c3c85A_ |
|
not modelled |
85.3 |
19 |
PDB header:transport protein
Chain: A: PDB Molecule:putative glutathione-regulated potassium-efflux system
PDBTitle: crystal structure of trka domain of putative glutathione-regulated2 potassium-efflux kefb from vibrio parahaemolyticus
|
| 97 | d1wdka3 |
|
not modelled |
85.1 |
24 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:6-phosphogluconate dehydrogenase-like, N-terminal domain |
| 98 | c3nt5B_ |
|
not modelled |
85.0 |
23 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:inositol 2-dehydrogenase/d-chiro-inositol 3-dehydrogenase;
PDBTitle: crystal structure of myo-inositol dehydrogenase from bacillus subtilis2 with bound cofactor and product inosose
|
| 99 | c2graA_ |
|
not modelled |
85.0 |
16 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:pyrroline-5-carboxylate reductase 1;
PDBTitle: crystal structure of human pyrroline-5-carboxylate reductase complexed2 with nadp
|
| 100 | c3dzbA_ |
|
not modelled |
84.8 |
17 |
PDB header:biosynthetic protein
Chain: A: PDB Molecule:prephenate dehydrogenase;
PDBTitle: crystal structure of prephenate dehydrogenase from streptococcus2 thermophilus
|
| 101 | c2pv7B_ |
|
not modelled |
84.4 |
23 |
PDB header:isomerase, oxidoreductase
Chain: B: PDB Molecule:t-protein [includes: chorismate mutase (ec 5.4.99.5) (cm)
PDBTitle: crystal structure of chorismate mutase / prephenate dehydrogenase2 (tyra) (1574749) from haemophilus influenzae rd at 2.00 a resolution
|
| 102 | c3e8xA_ |
|
not modelled |
84.3 |
22 |
PDB header:structural genomics, unknown function
Chain: A: PDB Molecule:putative nad-dependent epimerase/dehydratase;
PDBTitle: putative nad-dependent epimerase/dehydratase from bacillus halodurans.
|
| 103 | c2ggsB_ |
|
not modelled |
84.0 |
21 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:273aa long hypothetical dtdp-4-dehydrorhamnose
PDBTitle: crystal structure of hypothetical dtdp-4-dehydrorhamnose2 reductase from sulfolobus tokodaii
|
| 104 | d1s6ya1 |
|
not modelled |
84.0 |
16 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:LDH N-terminal domain-like |
| 105 | d1e5qa1 |
|
not modelled |
83.9 |
20 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:Glyceraldehyde-3-phosphate dehydrogenase-like, N-terminal domain |
| 106 | c3n7uD_ |
|
not modelled |
83.2 |
31 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:formate dehydrogenase;
PDBTitle: nad-dependent formate dehydrogenase from higher-plant arabidopsis2 thaliana in complex with nad and azide
|
| 107 | c2glxD_ |
|
not modelled |
83.2 |
14 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:1,5-anhydro-d-fructose reductase;
PDBTitle: crystal structure analysis of bacterial 1,5-af reductase
|
| 108 | c3e18A_ |
|
not modelled |
82.9 |
14 |
PDB header:oxidoreductase
Chain: A: PDB Molecule:oxidoreductase;
PDBTitle: crystal structure of nad-binding protein from listeria innocua
|
| 109 | c1yj8C_ |
|
not modelled |
82.9 |
20 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:glycerol-3-phosphate dehydrogenase;
PDBTitle: initial structural analysis of plasmodium falciparum glycerol-3-2 phosphate dehydrogenase
|
| 110 | c1xeaD_ |
|
not modelled |
82.9 |
28 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:oxidoreductase, gfo/idh/moca family;
PDBTitle: crystal structure of a gfo/idh/moca family oxidoreductase2 from vibrio cholerae
|
| 111 | c2q3eH_ |
|
not modelled |
82.8 |
32 |
PDB header:oxidoreductase
Chain: H: PDB Molecule:udp-glucose 6-dehydrogenase;
PDBTitle: structure of human udp-glucose dehydrogenase complexed with nadh and2 udp-glucose
|
| 112 | d1ojua1 |
|
not modelled |
82.5 |
17 |
Fold:NAD(P)-binding Rossmann-fold domains
Superfamily:NAD(P)-binding Rossmann-fold domains
Family:LDH N-terminal domain-like |
| 113 | c2ho3D_ |
|
not modelled |
82.1 |
21 |
PDB header:oxidoreductase
Chain: D: PDB Molecule:oxidoreductase, gfo/idh/moca family;
PDBTitle: crystal structure of oxidoreductase, gfo/idh/moca family from2 streptococcus pneumoniae
|
| 114 | c3gvpB_ |
|
not modelled |
81.9 |
19 |
PDB header:hydrolase
Chain: B: PDB Molecule:adenosylhomocysteinase 3;
PDBTitle: human sahh-like domain of human adenosylhomocysteinase 3
|
| 115 | c3e9mC_ |
|
not modelled |
81.9 |
12 |
PDB header:oxidoreductase
Chain: C: PDB Molecule:oxidoreductase, gfo/idh/moca family;
PDBTitle: crystal structure of an oxidoreductase from enterococcus2 faecalis
|
| 116 | c1vkzA_ |
|
not modelled |
81.8 |
31 |
PDB header:ligase
Chain: A: PDB Molecule:phosphoribosylamine--glycine ligase;
PDBTitle: crystal structure of phosphoribosylamine--glycine ligase (tm1250) from2 thermotoga maritima at 2.30 a resolution
|
| 117 | c2q4eB_ |
|
not modelled |
81.5 |
12 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:probable oxidoreductase at4g09670;
PDBTitle: ensemble refinement of the protein crystal structure of gene product2 from arabidopsis thaliana at4g09670
|
| 118 | c3dhyC_ |
|
not modelled |
81.3 |
31 |
PDB header:hydrolase
Chain: C: PDB Molecule:adenosylhomocysteinase;
PDBTitle: crystal structures of mycobacterium tuberculosis s-adenosyl-l-2 homocysteine hydrolase in ternary complex with substrate and3 inhibitors
|
| 119 | c2dt5A_ |
|
not modelled |
80.9 |
21 |
PDB header:dna binding protein
Chain: A: PDB Molecule:at-rich dna-binding protein;
PDBTitle: crystal structure of ttha1657 (at-rich dna-binding protein) from2 thermus thermophilus hb8
|
| 120 | c1d4fD_ |
|
not modelled |
80.7 |
18 |
PDB header:hydrolase
Chain: D: PDB Molecule:s-adenosylhomocysteine hydrolase;
PDBTitle: crystal structure of recombinant rat-liver d244e mutant s-2 adenosylhomocysteine hydrolase
|