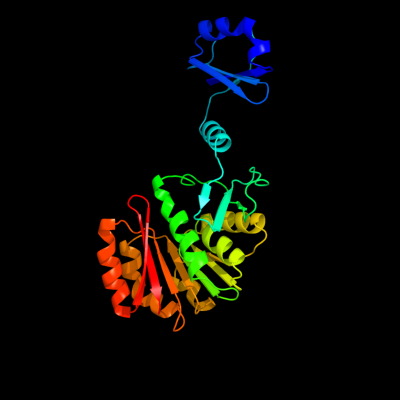
1 d2nxca1
100.0
34
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Ribosomal protein L11 methyltransferase PrmA2 c3grzA_
100.0
34
PDB header: transferaseChain: A: PDB Molecule: ribosomal protein l11 methyltransferase;PDBTitle: crystal structure of ribosomal protein l11 methylase from2 lactobacillus delbrueckii subsp. bulgaricus
3 c2yx1A_
99.9
13
PDB header: transferaseChain: A: PDB Molecule: hypothetical protein mj0883;PDBTitle: crystal structure of m.jannaschii trna m1g37 methyltransferase
4 c3m70A_
99.9
14
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: tellurite resistance protein tehb homolog;PDBTitle: crystal structure of tehb from haemophilus influenzae
5 d2b3ta1
99.9
23
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: N5-glutamine methyltransferase, HemK6 d1dusa_
99.9
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Hypothetical protein MJ08827 c3lcvB_
99.9
18
PDB header: transferaseChain: B: PDB Molecule: sisomicin-gentamicin resistance methylase sgm;PDBTitle: crystal structure of antibiotic related methyltransferase
8 c3e05B_
99.9
18
PDB header: transferaseChain: B: PDB Molecule: precorrin-6y c5,15-methyltransferase (decarboxylating);PDBTitle: crystal structure of precorrin-6y c5,15-methyltransferase from2 geobacter metallireducens gs-15
9 c2b78A_
99.9
16
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: hypothetical protein smu.776;PDBTitle: a putative sam-dependent methyltransferase from2 streptococcus mutans
10 d2frna1
99.9
19
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Met-10+ protein-like11 d2fhpa1
99.9
20
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: YhhF-like12 c3lecA_
99.9
15
PDB header: structure genomics, unknown functionChain: A: PDB Molecule: nadb-rossmann superfamily protein;PDBTitle: the crystal structure of a protein in the nadb-rossmann2 superfamily from streptococcus agalactiae to 1.8a
13 d1kpia_
99.9
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Mycolic acid cyclopropane synthase14 d1l3ia_
99.9
22
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Precorrin-6Y methyltransferase (CbiT)15 d1l1ea_
99.8
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Mycolic acid cyclopropane synthase16 d2as0a2
99.8
27
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: hypothetical RNA methyltransferase17 c3dmgA_
99.8
20
PDB header: transferaseChain: A: PDB Molecule: probable ribosomal rna small subunit methyltransferase;PDBTitle: t. thermophilus 16s rrna n2 g1207 methyltransferase (rsmc) in complex2 with adohcy
18 d2igta1
99.8
13
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: hypothetical RNA methyltransferase19 c3c0kB_
99.8
19
PDB header: transferaseChain: B: PDB Molecule: upf0064 protein yccw;PDBTitle: crystal structure of a ribosomal rna methyltranferase
20 d1nv8a_
99.8
26
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: N5-glutamine methyltransferase, HemK21 c2fk8A_
not modelled
99.8
15
PDB header: transferaseChain: A: PDB Molecule: methoxy mycolic acid synthase 4;PDBTitle: crystal structure of hma (mmaa4) from mycobacterium tuberculosis2 complexed with s-adenosylmethionine
22 c3njrB_
not modelled
99.8
21
PDB header: transferaseChain: B: PDB Molecule: precorrin-6y methylase;PDBTitle: crystal structure of c-terminal domain of precorrin-6y c5,15-2 methyltransferase from rhodobacter capsulatus
23 c3a26A_
not modelled
99.8
20
PDB header: transferaseChain: A: PDB Molecule: uncharacterized protein ph0793;PDBTitle: crystal structure of p. horikoshii tyw2 in complex with2 mesado
24 c3evzA_
not modelled
99.8
16
PDB header: transferaseChain: A: PDB Molecule: methyltransferase;PDBTitle: crystal strucure of methyltransferase from pyrococcus furiosus
25 c3ocjA_
not modelled
99.8
19
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: putative exported protein;PDBTitle: the crystal structure of a possilbe exported protein from bordetella2 parapertussis
26 c3ku1E_
not modelled
99.8
16
PDB header: transferaseChain: E: PDB Molecule: sam-dependent methyltransferase;PDBTitle: crystal structure of streptococcus pneumoniae sp1610, a2 putative trna (m1a22) methyltransferase, in complex with s-3 adenosyl-l-methionine
27 d1kpga_
not modelled
99.8
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Mycolic acid cyclopropane synthase28 d2fk8a1
not modelled
99.8
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Mycolic acid cyclopropane synthase29 d1tpya_
not modelled
99.8
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Mycolic acid cyclopropane synthase30 d2b78a2
not modelled
99.8
16
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: hypothetical RNA methyltransferase31 c3lpmA_
not modelled
99.8
16
PDB header: transferaseChain: A: PDB Molecule: putative methyltransferase;PDBTitle: crystal structure of putative methyltransferase small domain protein2 from listeria monocytogenes
32 c3fzgA_
not modelled
99.8
17
PDB header: transferaseChain: A: PDB Molecule: 16s rrna methylase;PDBTitle: structure of the 16s rrna methylase arma
33 d1wxxa2
not modelled
99.8
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: hypothetical RNA methyltransferase34 c3lccA_
not modelled
99.8
19
PDB header: transferaseChain: A: PDB Molecule: putative methyl chloride transferase;PDBTitle: structure of a sam-dependent halide methyltransferase from arabidopsis2 thaliana
35 c2as0A_
not modelled
99.8
26
PDB header: transferaseChain: A: PDB Molecule: hypothetical protein ph1915;PDBTitle: crystal structure of ph1915 (apc 5817): a hypothetical rna2 methyltransferase
36 c3gnlB_
not modelled
99.8
16
PDB header: structural genomics, unknown functionChain: B: PDB Molecule: uncharacterized protein, duf633, lmof2365_1472;PDBTitle: structure of uncharacterized protein (lmof2365_1472) from2 listeria monocytogenes serotype 4b
37 c3b89A_
not modelled
99.8
19
PDB header: transferaseChain: A: PDB Molecule: 16s rrna methylase;PDBTitle: crystal structure of rrna methylase from escherichia coli
38 d1yb2a1
not modelled
99.8
16
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: tRNA(1-methyladenosine) methyltransferase-like39 c1yb2A_
not modelled
99.8
16
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: hypothetical protein ta0852;PDBTitle: structure of a putative methyltransferase from thermoplasma2 acidophilum.
40 c3p9nA_
not modelled
99.8
20
PDB header: transferaseChain: A: PDB Molecule: possible methyltransferase (methylase);PDBTitle: rv2966c of m. tuberculosis is a rsmd-like methyltransferase
41 d2ifta1
not modelled
99.8
16
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: YhhF-like42 c2pjdA_
not modelled
99.8
20
PDB header: transferaseChain: A: PDB Molecule: ribosomal rna small subunit methyltransferase c;PDBTitle: crystal structure of 16s rrna methyltransferase rsmc
43 c3q87B_
not modelled
99.8
23
PDB header: transferase activator/transferaseChain: B: PDB Molecule: n6 adenine specific dna methylase;PDBTitle: structure of eukaryotic translation termination complex2 methyltransferase mtq2-trm112
44 d2fpoa1
not modelled
99.8
16
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: YhhF-like45 c3e7pA_
not modelled
99.8
21
PDB header: transferaseChain: A: PDB Molecule: putative methyltransferase;PDBTitle: crystal structure of of putative methyltransferase from bacteroides2 vulgatus atcc 8482
46 d1xtpa_
not modelled
99.8
10
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: AD-003 protein-like47 c3mtiA_
not modelled
99.8
16
PDB header: transferaseChain: A: PDB Molecule: rrna methylase;PDBTitle: the crystal structure of a rrna methylase from streptococcus2 thermophilus to 1.95a
48 d1o54a_
not modelled
99.8
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: tRNA(1-methyladenosine) methyltransferase-like49 c3hm2G_
not modelled
99.8
19
PDB header: transferaseChain: G: PDB Molecule: precorrin-6y c5,15-methyltransferase;PDBTitle: crystal structure of putative precorrin-6y c5,15-2 methyltransferase targeted domain from corynebacterium3 diphtheriae
50 c3opnA_
not modelled
99.8
18
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: putative hemolysin;PDBTitle: the crystal structure of a putative hemolysin from lactococcus lactis
51 d2ex4a1
not modelled
99.8
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: AD-003 protein-like52 c1z3cA_
not modelled
99.8
19
PDB header: transferaseChain: A: PDB Molecule: mrna capping enzyme;PDBTitle: encephalitozooan cuniculi mrna cap (guanine-n7)2 methyltransferasein complexed with azoadomet
53 d1yzha1
not modelled
99.8
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: TrmB-like54 c2pwyB_
not modelled
99.8
19
PDB header: transferaseChain: B: PDB Molecule: trna (adenine-n(1)-)-methyltransferase;PDBTitle: crystal structure of a m1a58 trna methyltransferase
55 d2a14a1
not modelled
99.8
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Arylamine N-methyltransferase56 d1ri5a_
not modelled
99.8
19
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: mRNA cap (Guanine N-7) methyltransferase57 c3dlcA_
not modelled
99.8
22
PDB header: transferaseChain: A: PDB Molecule: putative s-adenosyl-l-methionine-dependentPDBTitle: crystal structure of a putative s-adenosyl-l-methionine-dependent2 methyltransferase (mmp1179) from methanococcus maripaludis at 1.15 a3 resolution
58 c2ozvA_
not modelled
99.8
22
PDB header: transferaseChain: A: PDB Molecule: hypothetical protein atu0636;PDBTitle: crystal structure of a predicted o-methyltransferase, protein atu6362 from agrobacterium tumefaciens.
59 d1im8a_
not modelled
99.8
19
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Hypothetical protein HI0319 (YecO)60 c2yvlB_
not modelled
99.8
18
PDB header: transferaseChain: B: PDB Molecule: hypothetical protein;PDBTitle: crystal structure of trna (m1a58) methyltransferase trmi from aquifex2 aeolicus
61 c3e23A_
not modelled
99.8
25
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: uncharacterized protein rpa2492;PDBTitle: crystal structure of the rpa2492 protein in complex with2 sam from rhodopseudomonas palustris, northeast structural3 genomics consortium target rpr299
62 c3eeyJ_
not modelled
99.7
20
PDB header: transferaseChain: J: PDB Molecule: putative rrna methylase;PDBTitle: crystal structure of putative rrna-methylase from clostridium2 thermocellum
63 d1wy7a1
not modelled
99.7
19
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Ta1320-like64 c3f4kA_
not modelled
99.7
19
PDB header: transferaseChain: A: PDB Molecule: putative methyltransferase;PDBTitle: crystal structure of a probable methyltransferase from2 bacteroides thetaiotaomicron. northeast structural3 genomics target btr309.
65 c3a27A_
not modelled
99.7
13
PDB header: transferaseChain: A: PDB Molecule: uncharacterized protein mj1557;PDBTitle: crystal structure of m. jannaschii tyw2 in complex with2 adomet
66 c1wxwA_
not modelled
99.7
19
PDB header: transferaseChain: A: PDB Molecule: hypothetical protein ttha1280;PDBTitle: crystal structure of tt1595, a putative sam-dependent2 methyltransferase from thermus thermophillus hb8
67 c2iipD_
not modelled
99.7
22
PDB header: transferaseChain: D: PDB Molecule: nicotinamide n-methyltransferase;PDBTitle: human nicotinamide n-methyltransferase
68 c3h2bB_
not modelled
99.7
12
PDB header: transferaseChain: B: PDB Molecule: sam-dependent methyltransferase;PDBTitle: crystal structure of the sam-dependent methyltransferase2 cg3271 from corynebacterium glutamicum in complex with s-3 adenosyl-l-homocysteine and pyrophosphate. northeast4 structural genomics consortium target cgr113a
69 d1zx0a1
not modelled
99.7
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Guanidinoacetate methyltransferase70 c2yxdA_
not modelled
99.7
14
PDB header: transferaseChain: A: PDB Molecule: probable cobalt-precorrin-6y c(15)-methyltransferasePDBTitle: crystal structure of cobalamin biosynthesis precorrin 8w decarboxylase2 (cbit)
71 c3ujcA_
not modelled
99.7
18
PDB header: transferaseChain: A: PDB Molecule: phosphoethanolamine n-methyltransferase;PDBTitle: phosphoethanolamine methyltransferase mutant (h132a) from plasmodium2 falciparum in complex with phosphocholine
72 d2fcaa1
not modelled
99.7
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: TrmB-like73 d1xxla_
not modelled
99.7
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: UbiE/COQ5-like74 d2o57a1
not modelled
99.7
12
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Mycolic acid cyclopropane synthase75 d1p1ca_
not modelled
99.7
14
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Guanidinoacetate methyltransferase76 c3mb5A_
not modelled
99.7
17
PDB header: transferaseChain: A: PDB Molecule: sam-dependent methyltransferase;PDBTitle: crystal structure of p. abyssi trna m1a58 methyltransferase in complex2 with s-adenosyl-l-methionine
77 c2pxxA_
not modelled
99.7
16
PDB header: transferaseChain: A: PDB Molecule: uncharacterized protein mgc2408;PDBTitle: human putative methyltransferase mgc2408
78 d1nt2a_
not modelled
99.7
12
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Fibrillarin homologue79 c1vl5B_
not modelled
99.7
18
PDB header: transferaseChain: B: PDB Molecule: unknown conserved protein bh2331;PDBTitle: crystal structure of a putative methyltransferase (bh2331) from2 bacillus halodurans c-125 at 1.95 a resolution
80 d1xcla_
not modelled
99.7
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Guanidinoacetate methyltransferase81 d1vl5a_
not modelled
99.7
18
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: UbiE/COQ5-like82 d1xdza_
not modelled
99.7
18
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Glucose-inhibited division protein B (GidB)83 d1ve3a1
not modelled
99.7
18
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: CAC2371-like84 c3b3jA_
not modelled
99.7
22
PDB header: transferaseChain: A: PDB Molecule: histone-arginine methyltransferase carm1;PDBTitle: the 2.55 a crystal structure of the apo catalytic domain of2 coactivator-associated arginine methyl transferase i(carm1:28-507,3 residues 28-146 and 479-507 not ordered)
85 d1g8aa_
not modelled
99.7
18
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Fibrillarin homologue86 d1g6q1_
not modelled
99.7
29
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Arginine methyltransferase87 d1f3la_
not modelled
99.7
26
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Arginine methyltransferase88 c3g5tA_
not modelled
99.7
17
PDB header: transferaseChain: A: PDB Molecule: trans-aconitate 3-methyltransferase;PDBTitle: crystal structure of trans-aconitate 3-methyltransferase2 from yeast
89 c2esrB_
not modelled
99.7
15
PDB header: transferaseChain: B: PDB Molecule: methyltransferase;PDBTitle: conserved hypothetical protein- streptococcus pyogenes
90 c3merA_
not modelled
99.7
18
PDB header: structural genomics, unknown functionChain: A: PDB Molecule: slr1183 protein;PDBTitle: crystal structure of the methyltransferase slr1183 from2 synechocystis sp. pcc 6803, northeast structural genomics3 consortium target sgr145
91 d2fyta1
not modelled
99.7
25
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Arginine methyltransferase92 d1oria_
not modelled
99.7
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Arginine methyltransferase93 c3bgvC_
not modelled
99.7
19
PDB header: transferaseChain: C: PDB Molecule: mrna cap guanine-n7 methyltransferase;PDBTitle: crystal structure of mrna cap guanine-n7 methyltransferase2 in complex with sah
94 d1uwva2
not modelled
99.7
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: (Uracil-5-)-methyltransferase95 d1nkva_
not modelled
99.7
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Hypothetical Protein YjhP96 d1ws6a1
not modelled
99.7
24
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: YhhF-like97 d2h00a1
not modelled
99.7
16
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Methyltransferase 10 domain98 d2esra1
not modelled
99.7
20
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: YhhF-like99 c3dxyA_
not modelled
99.7
15
PDB header: transferaseChain: A: PDB Molecule: trna (guanine-n(7)-)-methyltransferase;PDBTitle: crystal structure of ectrmb in complex with sam
100 c1orhA_
not modelled
99.7
21
PDB header: transferaseChain: A: PDB Molecule: protein arginine n-methyltransferase 1;PDBTitle: structure of the predominant protein arginine methyltransferase prmt1
101 c3g07C_
not modelled
99.7
28
PDB header: transferaseChain: C: PDB Molecule: 7sk snrna methylphosphate capping enzyme;PDBTitle: methyltransferase domain of human bicoid-interacting protein2 3 homolog (drosophila)
102 c3cggB_
not modelled
99.7
20
PDB header: transferaseChain: B: PDB Molecule: sam-dependent methyltransferase;PDBTitle: crystal structure of tehb-like sam-dependent methyltransferase2 (np_600671.1) from corynebacterium glutamicum atcc 13032 kitasato at3 2.00 a resolution
103 c3bkxB_
not modelled
99.7
18
PDB header: transferaseChain: B: PDB Molecule: sam-dependent methyltransferase;PDBTitle: crystal structure of cyclopropane-fatty-acyl-phospholipid synthase-2 like protein (yp_807781.1) from lactobacillus casei atcc 334 at 1.853 a resolution
104 c3ofkA_
not modelled
99.7
16
PDB header: transferaseChain: A: PDB Molecule: nodulation protein s;PDBTitle: crystal structure of n-methyltransferase nods from bradyrhizobium2 japonicum wm9 in complex with s-adenosyl-l-homocysteine (sah)
105 c3id5F_
not modelled
99.7
14
PDB header: transferase/ribosomal protein/rnaChain: F: PDB Molecule: fibrillarin-like rrna/trna 2'-o-methyltransferase;PDBTitle: crystal structure of sulfolobus solfataricus c/d rnp assembled with2 nop5, fibrillarin, l7ae and a split half c/d rna
106 d1r74a_
not modelled
99.7
21
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Glycine N-methyltransferase107 d1jsxa_
not modelled
99.7
18
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Glucose-inhibited division protein B (GidB)108 d1xvaa_
not modelled
99.7
23
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Glycine N-methyltransferase109 d2gh1a1
not modelled
99.7
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: BC2162-like110 d2bzga1
not modelled
99.7
15
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Thiopurine S-methyltransferase111 c2p8jA_
not modelled
99.7
16
PDB header: transferaseChain: A: PDB Molecule: s-adenosylmethionine-dependent methyltransferase;PDBTitle: crystal structure of s-adenosylmethionine-dependent methyltransferase2 (np_349143.1) from clostridium acetobutylicum at 2.00 a resolution
112 d1i9ga_
not modelled
99.7
23
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: tRNA(1-methyladenosine) methyltransferase-like113 d1prya_
not modelled
99.7
17
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Fibrillarin homologue114 c3busB_
not modelled
99.7
17
PDB header: transferaseChain: B: PDB Molecule: methyltransferase;PDBTitle: crystal structure of rebm
115 c2yr0A_
not modelled
99.7
14
PDB header: transferaseChain: A: PDB Molecule: hypothetical protein ttha0223;PDBTitle: crystal structure of hypothetical methyltransferase ttha0223 from2 thermus thermophilus hb8
116 c3axtA_
not modelled
99.7
22
PDB header: transferaseChain: A: PDB Molecule: probable n(2),n(2)-dimethylguanosine trna methyltransferasePDBTitle: complex structure of trna methyltransferase trm1 from aquifex aeolicus2 with s-adenosyl-l-methionine
117 d2g72a1
not modelled
99.7
20
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Arylamine N-methyltransferase118 d1i1na_
not modelled
99.7
25
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Protein-L-isoaspartyl O-methyltransferase119 d1jg1a_
not modelled
99.7
20
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Protein-L-isoaspartyl O-methyltransferase120 d1vbfa_
not modelled
99.7
19
Fold: S-adenosyl-L-methionine-dependent methyltransferasesSuperfamily: S-adenosyl-L-methionine-dependent methyltransferasesFamily: Protein-L-isoaspartyl O-methyltransferase