| 1 | c2pfuA_ |
|
 |
99.8 |
100 |
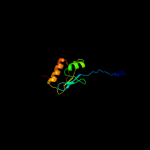
PDB header:transport protein
Chain: A: PDB Molecule:biopolymer transport exbd protein;
PDBTitle: nmr strcuture determination of the periplasmic domain of exbd from2 e.coli
|
| 2 | c2jwlB_ |
|
 |
99.4 |
28 |
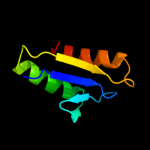
PDB header:membrane protein
Chain: B: PDB Molecule:protein tolr;
PDBTitle: solution structure of periplasmic domain of tolr from h.2 influenzae with saxs data
|
| 3 | d2axtk1 |
|
 |
49.0 |
31 |
Fold:Single transmembrane helix
Superfamily:Photosystem II reaction center protein K, PsbK
Family:PsbK-like |
| 4 | c3a0bK_ |
|
 |
45.2 |
31 |
PDB header:electron transport
Chain: K: PDB Molecule:photosystem ii reaction center protein k;
PDBTitle: crystal structure of br-substituted photosystem ii complex
|
| 5 | c3e6eC_ |
|
 |
35.6 |
10 |
PDB header:isomerase
Chain: C: PDB Molecule:alanine racemase;
PDBTitle: crystal structure of alanine racemase from e.faecalis2 complex with cycloserine
|
| 6 | c3a0bk_ |
|
 |
35.2 |
31 |
PDB header:electron transport
Chain: K: PDB Molecule:photosystem ii reaction center protein k;
PDBTitle: crystal structure of br-substituted photosystem ii complex
|
| 7 | d1bd0a2 |
|
 |
29.5 |
19 |
Fold:TIM beta/alpha-barrel
Superfamily:PLP-binding barrel
Family:Alanine racemase-like, N-terminal domain |
| 8 | c2jvfA_ |
|
 |
28.2 |
21 |
PDB header:de novo protein
Chain: A: PDB Molecule:de novo protein m7;
PDBTitle: solution structure of m7, a computationally-designed2 artificial protein
|
| 9 | c2re3A_ |
|
 |
27.6 |
13 |
PDB header:unknown function
Chain: A: PDB Molecule:uncharacterized protein;
PDBTitle: crystal structure of a duf1285 family protein (spo_0140) from2 silicibacter pomeroyi dss-3 at 2.50 a resolution
|
| 10 | c3a0hk_ |
|
 |
26.0 |
31 |
PDB header:electron transport
Chain: K: PDB Molecule:photosystem ii reaction center protein k;
PDBTitle: crystal structure of i-substituted photosystem ii complex
|
| 11 | d2e8aa1 |
|
 |
25.1 |
18 |
Fold:Ribonuclease H-like motif
Superfamily:Actin-like ATPase domain
Family:Actin/HSP70 |
| 12 | d1jcea1 |
|
 |
25.1 |
16 |
Fold:Ribonuclease H-like motif
Superfamily:Actin-like ATPase domain
Family:Actin/HSP70 |
| 13 | c1qysA_ |
|
 |
22.5 |
17 |
PDB header:de novo protein
Chain: A: PDB Molecule:top7;
PDBTitle: crystal structure of top7: a computationally designed2 protein with a novel fold
|
| 14 | d1dkgd1 |
|
 |
20.9 |
14 |
Fold:Ribonuclease H-like motif
Superfamily:Actin-like ATPase domain
Family:Actin/HSP70 |
| 15 | d1bupa1 |
|
 |
19.8 |
18 |
Fold:Ribonuclease H-like motif
Superfamily:Actin-like ATPase domain
Family:Actin/HSP70 |
| 16 | d1vfsa2 |
|
 |
19.8 |
16 |
Fold:TIM beta/alpha-barrel
Superfamily:PLP-binding barrel
Family:Alanine racemase-like, N-terminal domain |
| 17 | c1ur8B_ |
|
 |
18.4 |
12 |
PDB header:hydrolase
Chain: B: PDB Molecule:chitinase b;
PDBTitle: interactions of a family 18 chitinase with the designed2 inhibitor hm508, and its degradation product,3 chitobiono-delta-lactone
|
| 18 | c3ugsB_ |
|
 |
18.3 |
11 |
PDB header:transferase
Chain: B: PDB Molecule:undecaprenyl pyrophosphate synthase;
PDBTitle: crystal structure of a probable undecaprenyl diphosphate synthase2 (upps) from campylobacter jejuni
|
| 19 | c1vftA_ |
|
 |
17.8 |
16 |
PDB header:isomerase
Chain: A: PDB Molecule:alanine racemase;
PDBTitle: crystal structure of l-cycloserine-bound form of alanine2 racemase from d-cycloserine-producing streptomyces3 lavendulae
|
| 20 | c3ghfA_ |
|
 |
17.5 |
11 |
PDB header:cell cycle
Chain: A: PDB Molecule:septum site-determining protein minc;
PDBTitle: crystal structure of the septum site-determining protein2 minc from salmonella typhimurium
|
| 21 | c3co8B_ |
|
not modelled |
16.6 |
12 |
PDB header:isomerase
Chain: B: PDB Molecule:alanine racemase;
PDBTitle: crystal structure of alanine racemase from oenococcus oeni
|
| 22 | c3lupA_ |
|
not modelled |
16.5 |
18 |
PDB header:structure genomics, unknown function
Chain: A: PDB Molecule:degv family protein;
PDBTitle: crystal structure of fatty acid binding degv family protein sag13422 from streptococcus agalactiae
|
| 23 | c2c2xB_ |
|
not modelled |
16.3 |
9 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:methylenetetrahydrofolate dehydrogenase-
PDBTitle: three dimensional structure of bifunctional2 methylenetetrahydrofolate dehydrogenase-cyclohydrolase3 from mycobacterium tuberculosis
|
| 24 | d1edza2 |
|
not modelled |
16.1 |
9 |
Fold:Aminoacid dehydrogenase-like, N-terminal domain
Superfamily:Aminoacid dehydrogenase-like, N-terminal domain
Family:Tetrahydrofolate dehydrogenase/cyclohydrolase |
| 25 | c3me8B_ |
|
not modelled |
15.8 |
11 |
PDB header:electron transport
Chain: B: PDB Molecule:putative uncharacterized protein;
PDBTitle: crystal structure of putative electron transfer protein aq_2194 from2 aquifex aeolicus vf5
|
| 26 | c1niuA_ |
|
not modelled |
15.7 |
19 |
PDB header:isomerase
Chain: A: PDB Molecule:alanine racemase;
PDBTitle: alanine racemase with bound inhibitor derived from l-2 cycloserine
|
| 27 | d1qyia_ |
|
not modelled |
14.9 |
15 |
Fold:HAD-like
Superfamily:HAD-like
Family:Hypothetical protein MW1667 (SA1546) |
| 28 | d1goia2 |
|
not modelled |
14.8 |
13 |
Fold:TIM beta/alpha-barrel
Superfamily:(Trans)glycosidases
Family:Type II chitinase |
| 29 | c3oo2A_ |
|
not modelled |
13.6 |
12 |
PDB header:isomerase
Chain: A: PDB Molecule:alanine racemase 1;
PDBTitle: 2.37 angstrom resolution crystal structure of an alanine racemase2 (alr) from staphylococcus aureus subsp. aureus col
|
| 30 | d2d0oa1 |
|
not modelled |
12.5 |
17 |
Fold:The "swivelling" beta/beta/alpha domain
Superfamily:Swiveling domain of dehydratase reactivase alpha subunit
Family:Swiveling domain of dehydratase reactivase alpha subunit |
| 31 | c3pm6B_ |
|
not modelled |
12.3 |
14 |
PDB header:lyase
Chain: B: PDB Molecule:putative fructose-bisphosphate aldolase;
PDBTitle: crystal structure of a putative fructose-1,6-biphosphate aldolase from2 coccidioides immitis solved by combined sad mr
|
| 32 | c3outC_ |
|
not modelled |
11.9 |
15 |
PDB header:isomerase
Chain: C: PDB Molecule:glutamate racemase;
PDBTitle: crystal structure of glutamate racemase from francisella tularensis2 subsp. tularensis schu s4 in complex with d-glutamate.
|
| 33 | d1a9xa4 |
|
not modelled |
11.7 |
11 |
Fold:PreATP-grasp domain
Superfamily:PreATP-grasp domain
Family:BC N-terminal domain-like |
| 34 | d2j9ga2 |
|
not modelled |
11.5 |
7 |
Fold:PreATP-grasp domain
Superfamily:PreATP-grasp domain
Family:BC N-terminal domain-like |
| 35 | c2g7zB_ |
|
not modelled |
11.4 |
17 |
PDB header:structural genomics, unknown function
Chain: B: PDB Molecule:conserved hypothetical protein spy1493;
PDBTitle: conserved degv-like protein of unknown function from streptococcus2 pyogenes m1 gas binds long-chain fatty acids
|
| 36 | c3iprC_ |
|
not modelled |
10.8 |
17 |
PDB header:transferase
Chain: C: PDB Molecule:pts system, iia component;
PDBTitle: crystal structure of the enterococcus faecalis gluconate2 specific eiia phosphotransferase system component
|
| 37 | d1b0aa2 |
|
not modelled |
10.8 |
4 |
Fold:Aminoacid dehydrogenase-like, N-terminal domain
Superfamily:Aminoacid dehydrogenase-like, N-terminal domain
Family:Tetrahydrofolate dehydrogenase/cyclohydrolase |
| 38 | d1wd5a_ |
|
not modelled |
10.8 |
11 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 39 | c1xfcB_ |
|
not modelled |
10.7 |
19 |
PDB header:isomerase
Chain: B: PDB Molecule:alanine racemase;
PDBTitle: the 1.9 a crystal structure of alanine racemase from mycobacterium2 tuberculosis contains a conserved entryway into the active site
|
| 40 | d2axth1 |
|
not modelled |
10.5 |
18 |
Fold:Single transmembrane helix
Superfamily:Photosystem II 10 kDa phosphoprotein PsbH
Family:PsbH-like |
| 41 | d1ne9a1 |
|
not modelled |
10.5 |
18 |
Fold:Acyl-CoA N-acyltransferases (Nat)
Superfamily:Acyl-CoA N-acyltransferases (Nat)
Family:FemXAB nonribosomal peptidyltransferases |
| 42 | d2p10a1 |
|
not modelled |
10.0 |
8 |
Fold:TIM beta/alpha-barrel
Superfamily:Phosphoenolpyruvate/pyruvate domain
Family:Mll9387-like |
| 43 | c2dy3B_ |
|
not modelled |
9.9 |
14 |
PDB header:isomerase
Chain: B: PDB Molecule:alanine racemase;
PDBTitle: crystal structure of alanine racemase from corynebacterium glutamicum
|
| 44 | c2ra9A_ |
|
not modelled |
9.5 |
5 |
PDB header:unknown function
Chain: A: PDB Molecule:uncharacterized protein duf1285;
PDBTitle: crystal structure of a duf1285 family protein (sbal_2486) from2 shewanella baltica os155 at 1.40 a resolution
|
| 45 | c2odoC_ |
|
not modelled |
9.4 |
15 |
PDB header:isomerase
Chain: C: PDB Molecule:alanine racemase;
PDBTitle: crystal structure of pseudomonas fluorescens alanine racemase
|
| 46 | d1f75a_ |
|
not modelled |
9.2 |
19 |
Fold:Undecaprenyl diphosphate synthase
Superfamily:Undecaprenyl diphosphate synthase
Family:Undecaprenyl diphosphate synthase |
| 47 | c1p74B_ |
|
not modelled |
9.2 |
21 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:shikimate 5-dehydrogenase;
PDBTitle: crystal structure of shikimate dehydrogenase (aroe) from2 haemophilus influenzae
|
| 48 | c3a0hY_ |
|
not modelled |
8.7 |
10 |
PDB header:electron transport
Chain: Y: PDB Molecule:photosystem ii reaction center protein ycf12;
PDBTitle: crystal structure of i-substituted photosystem ii complex
|
| 49 | c3bz1y_ |
|
not modelled |
8.7 |
10 |
PDB header:electron transport
Chain: Y: PDB Molecule:photosystem ii protein y;
PDBTitle: crystal structure of cyanobacterial photosystem ii (part 12 of 2). this file contains first monomer of psii dimer
|
| 50 | c3a0hy_ |
|
not modelled |
8.7 |
10 |
PDB header:electron transport
Chain: Y: PDB Molecule:photosystem ii reaction center protein ycf12;
PDBTitle: crystal structure of i-substituted photosystem ii complex
|
| 51 | c3a0by_ |
|
not modelled |
8.5 |
10 |
PDB header:electron transport
Chain: Y: PDB Molecule:photosystem ii reaction center protein ycf12;
PDBTitle: crystal structure of br-substituted photosystem ii complex
|
| 52 | d1gph11 |
|
not modelled |
8.4 |
15 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 53 | c3arcY_ |
|
not modelled |
8.3 |
10 |
PDB header:electron transport, photosynthesis
Chain: Y: PDB Molecule:protein ycf12;
PDBTitle: crystal structure of oxygen-evolving photosystem ii at 1.9 angstrom2 resolution
|
| 54 | c3arcy_ |
|
not modelled |
8.3 |
10 |
PDB header:electron transport, photosynthesis
Chain: Y: PDB Molecule:protein ycf12;
PDBTitle: crystal structure of oxygen-evolving photosystem ii at 1.9 angstrom2 resolution
|
| 55 | d1gvfa_ |
|
not modelled |
8.3 |
19 |
Fold:TIM beta/alpha-barrel
Superfamily:Aldolase
Family:Class II FBP aldolase |
| 56 | c1jp3A_ |
|
not modelled |
8.0 |
10 |
PDB header:transferase
Chain: A: PDB Molecule:undecaprenyl pyrophosphate synthase;
PDBTitle: structure of e.coli undecaprenyl pyrophosphate synthase
|
| 57 | d1ecfa1 |
|
not modelled |
7.9 |
20 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 58 | c3eooL_ |
|
not modelled |
7.8 |
15 |
PDB header:lyase
Chain: L: PDB Molecule:methylisocitrate lyase;
PDBTitle: 2.9a crystal structure of methyl-isocitrate lyase from2 burkholderia pseudomallei
|
| 59 | d2nn6d2 |
|
not modelled |
7.7 |
8 |
Fold:Ribonuclease PH domain 2-like
Superfamily:Ribonuclease PH domain 2-like
Family:Ribonuclease PH domain 2-like |
| 60 | d1hgxa_ |
|
not modelled |
7.6 |
8 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 61 | d1rvga_ |
|
not modelled |
7.5 |
13 |
Fold:TIM beta/alpha-barrel
Superfamily:Aldolase
Family:Class II FBP aldolase |
| 62 | c3qm3C_ |
|
not modelled |
7.4 |
9 |
PDB header:lyase
Chain: C: PDB Molecule:fructose-bisphosphate aldolase;
PDBTitle: 1.85 angstrom resolution crystal structure of fructose-bisphosphate2 aldolase (fba) from campylobacter jejuni
|
| 63 | c3gmgB_ |
|
not modelled |
7.3 |
26 |
PDB header:structural genomics, unknown function
Chain: B: PDB Molecule:uncharacterized protein rv1825/mt1873;
PDBTitle: crystal structure of an uncharacterized conserved protein2 from mycobacterium tuberculosis
|
| 64 | c3o8qB_ |
|
not modelled |
7.2 |
21 |
PDB header:oxidoreductase
Chain: B: PDB Molecule:shikimate 5-dehydrogenase i alpha;
PDBTitle: 1.45 angstrom resolution crystal structure of shikimate 5-2 dehydrogenase (aroe) from vibrio cholerae
|
| 65 | c2vfwB_ |
|
not modelled |
6.8 |
14 |
PDB header:transferase
Chain: B: PDB Molecule:short-chain z-isoprenyl diphosphate synthetase;
PDBTitle: rv1086 native
|
| 66 | d1ueha_ |
|
not modelled |
6.8 |
10 |
Fold:Undecaprenyl diphosphate synthase
Superfamily:Undecaprenyl diphosphate synthase
Family:Undecaprenyl diphosphate synthase |
| 67 | d1zhva2 |
|
not modelled |
6.7 |
37 |
Fold:Ferredoxin-like
Superfamily:ACT-like
Family:Atu0741-like |
| 68 | c3gycB_ |
|
not modelled |
6.7 |
32 |
PDB header:hydrolase
Chain: B: PDB Molecule:putative glycoside hydrolase;
PDBTitle: crystal structure of putative glycoside hydrolase (yp_001304622.1)2 from parabacteroides distasonis atcc 8503 at 1.85 a resolution
|
| 69 | c3a0bY_ |
|
not modelled |
6.7 |
10 |
PDB header:electron transport
Chain: Y: PDB Molecule:photosystem ii reaction center protein ycf12;
PDBTitle: crystal structure of br-substituted photosystem ii complex
|
| 70 | d1g8fa3 |
|
not modelled |
6.7 |
17 |
Fold:P-loop containing nucleoside triphosphate hydrolases
Superfamily:P-loop containing nucleoside triphosphate hydrolases
Family:ATP sulfurylase C-terminal domain |
| 71 | c2d2rA_ |
|
not modelled |
6.2 |
14 |
PDB header:transferase
Chain: A: PDB Molecule:undecaprenyl pyrophosphate synthase;
PDBTitle: crystal structure of helicobacter pylori undecaprenyl pyrophosphate2 synthase
|
| 72 | c2f40A_ |
|
not modelled |
6.0 |
18 |
PDB header:structural genomics, unknown function
Chain: A: PDB Molecule:hypothetical protein pf1455;
PDBTitle: structure of a novel protein from backbone-centered nmr data and nmr-2 assisted structure prediction
|
| 73 | d1i5ea_ |
|
not modelled |
5.8 |
11 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 74 | c2qlxA_ |
|
not modelled |
5.8 |
31 |
PDB header:isomerase
Chain: A: PDB Molecule:l-rhamnose mutarotase;
PDBTitle: crystal structure of rhamnose mutarotase rhau of rhizobium2 leguminosarum in complex with l-rhamnose
|
| 75 | c2qlwA_ |
|
not modelled |
5.8 |
31 |
PDB header:isomerase
Chain: A: PDB Molecule:rhau;
PDBTitle: crystal structure of rhamnose mutarotase rhau of rhizobium2 leguminosarum
|
| 76 | c2dcqA_ |
|
not modelled |
5.8 |
9 |
PDB header:unknown function
Chain: A: PDB Molecule:putative protein at4g01050;
PDBTitle: fully automated nmr structure determination of the2 rhodanese homology domain at4g01050(175-295) from3 arabidopsis thaliana
|
| 77 | c3oo2B_ |
|
not modelled |
5.7 |
16 |
PDB header:isomerase
Chain: B: PDB Molecule:alanine racemase 1;
PDBTitle: 2.37 angstrom resolution crystal structure of an alanine racemase2 (alr) from staphylococcus aureus subsp. aureus col
|
| 78 | d1zzoa1 |
|
not modelled |
5.7 |
16 |
Fold:Thioredoxin fold
Superfamily:Thioredoxin-like
Family:Glutathione peroxidase-like |
| 79 | d2bz1a1 |
|
not modelled |
5.5 |
20 |
Fold:RibA-like
Superfamily:RibA-like
Family:RibA-like |
| 80 | d1x8da1 |
|
not modelled |
5.5 |
13 |
Fold:Ferredoxin-like
Superfamily:Dimeric alpha+beta barrel
Family:YiiL-like |
| 81 | c1ecjB_ |
|
not modelled |
5.3 |
20 |
PDB header:transferase
Chain: B: PDB Molecule:glutamine phosphoribosylpyrophosphate
PDBTitle: escherichia coli glutamine phosphoribosylpyrophosphate2 (prpp) amidotransferase complexed with 2 amp per tetramer
|
| 82 | d1g9sa_ |
|
not modelled |
5.2 |
16 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 83 | c1yfzA_ |
|
not modelled |
5.2 |
14 |
PDB header:transferase
Chain: A: PDB Molecule:hypoxanthine-guanine phosphoribosyltransferase;
PDBTitle: novel imp binding in feedback inhibition of hypoxanthine-guanine2 phosphoribosyltransferase from thermoanaerobacter tengcongensis
|
| 84 | d1yfza1 |
|
not modelled |
5.2 |
14 |
Fold:PRTase-like
Superfamily:PRTase-like
Family:Phosphoribosyltransferases (PRTases) |
| 85 | c3l8cA_ |
|
not modelled |
5.2 |
15 |
PDB header:ligase
Chain: A: PDB Molecule:d-alanine--poly(phosphoribitol) ligase subunit 1;
PDBTitle: structure of probable d-alanine--poly(phosphoribitol) ligase2 subunit-1 from streptococcus pyogenes
|
| 86 | c3q94B_ |
|
not modelled |
5.2 |
12 |
PDB header:lyase
Chain: B: PDB Molecule:fructose-bisphosphate aldolase, class ii;
PDBTitle: the crystal structure of fructose 1,6-bisphosphate aldolase from2 bacillus anthracis str. 'ames ancestor'
|
| 87 | c3navB_ |
|
not modelled |
5.1 |
15 |
PDB header:lyase
Chain: B: PDB Molecule:tryptophan synthase alpha chain;
PDBTitle: crystal structure of an alpha subunit of tryptophan synthase from2 vibrio cholerae o1 biovar el tor str. n16961
|
| 88 | d1xd7a_ |
|
not modelled |
5.1 |
14 |
Fold:DNA/RNA-binding 3-helical bundle
Superfamily:"Winged helix" DNA-binding domain
Family:Transcriptional regulator Rrf2 |
| 89 | d1aopa1 |
|
not modelled |
5.1 |
15 |
Fold:Ferredoxin-like
Superfamily:Nitrite/Sulfite reductase N-terminal domain-like
Family:Duplicated SiR/NiR-like domains 1 and 3 |
| 90 | d1ujpa_ |
|
not modelled |
5.1 |
9 |
Fold:TIM beta/alpha-barrel
Superfamily:Ribulose-phoshate binding barrel
Family:Tryptophan biosynthesis enzymes |
| 91 | d1ulza2 |
|
not modelled |
5.1 |
14 |
Fold:PreATP-grasp domain
Superfamily:PreATP-grasp domain
Family:BC N-terminal domain-like |
| 92 | d1rhsa1 |
|
not modelled |
5.1 |
10 |
Fold:Rhodanese/Cell cycle control phosphatase
Superfamily:Rhodanese/Cell cycle control phosphatase
Family:Multidomain sulfurtransferase (rhodanese) |
| 93 | d1zj8a1 |
|
not modelled |
5.0 |
14 |
Fold:Ferredoxin-like
Superfamily:Nitrite/Sulfite reductase N-terminal domain-like
Family:Duplicated SiR/NiR-like domains 1 and 3 |

