| 1 |
|
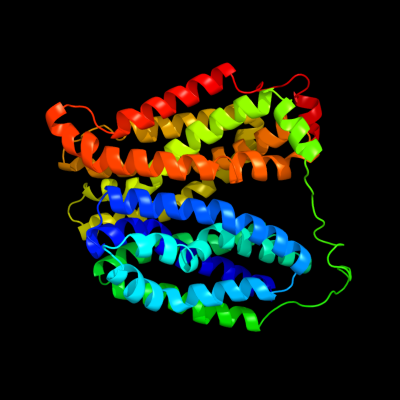
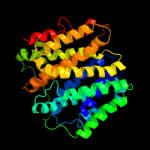
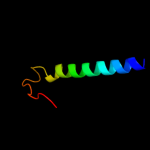
PDB 1pv7 chain A
Region: 2 - 417
Aligned: 406
Modelled: 410
Confidence: 100.0%
Identity: 10%
Fold: MFS general substrate transporter
Superfamily: MFS general substrate transporter
Family: LacY-like proton/sugar symporter
Phyre2
| 2 |
|
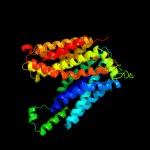
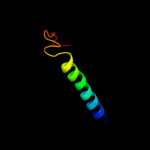
PDB 1pw4 chain A
Region: 2 - 416
Aligned: 406
Modelled: 409
Confidence: 100.0%
Identity: 11%
Fold: MFS general substrate transporter
Superfamily: MFS general substrate transporter
Family: Glycerol-3-phosphate transporter
Phyre2
| 3 |
|
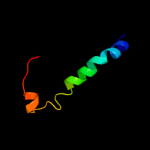
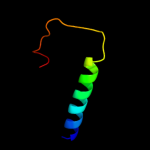
PDB 2xut chain C
Region: 7 - 398
Aligned: 391
Modelled: 392
Confidence: 100.0%
Identity: 14%
PDB header:transport protein
Chain: C: PDB Molecule:proton/peptide symporter family protein;
PDBTitle: crystal structure of a proton dependent oligopeptide (pot)2 family transporter.
Phyre2
| 4 |
|
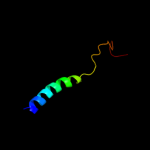
PDB 2gfp chain A
Region: 7 - 396
Aligned: 366
Modelled: 388
Confidence: 100.0%
Identity: 10%
PDB header:membrane protein
Chain: A: PDB Molecule:multidrug resistance protein d;
PDBTitle: structure of the multidrug transporter emrd from2 escherichia coli
Phyre2
| 5 |
|
PDB 3o7p chain A
Region: 8 - 405
Aligned: 388
Modelled: 398
Confidence: 100.0%
Identity: 9%
PDB header:transport protein
Chain: A: PDB Molecule:l-fucose-proton symporter;
PDBTitle: crystal structure of the e.coli fucose:proton symporter, fucp (n162a)
Phyre2
| 6 |
|
PDB 1fft chain B domain 2
Region: 380 - 418
Aligned: 39
Modelled: 39
Confidence: 50.9%
Identity: 15%
Fold: Transmembrane helix hairpin
Superfamily: Cytochrome c oxidase subunit II-like, transmembrane region
Family: Cytochrome c oxidase subunit II-like, transmembrane region
Phyre2
| 7 |
|
PDB 2knc chain A
Region: 380 - 417
Aligned: 38
Modelled: 38
Confidence: 42.1%
Identity: 11%
PDB header:cell adhesion
Chain: A: PDB Molecule:integrin alpha-iib;
PDBTitle: platelet integrin alfaiib-beta3 transmembrane-cytoplasmic2 heterocomplex
Phyre2
| 8 |
|
PDB 2rdd chain B
Region: 390 - 417
Aligned: 28
Modelled: 28
Confidence: 29.3%
Identity: 7%
PDB header:membrane protein/transport protein
Chain: B: PDB Molecule:upf0092 membrane protein yajc;
PDBTitle: x-ray crystal structure of acrb in complex with a novel2 transmembrane helix.
Phyre2
| 9 |
|
PDB 3dtu chain B domain 2
Region: 382 - 418
Aligned: 36
Modelled: 37
Confidence: 22.2%
Identity: 17%
Fold: Transmembrane helix hairpin
Superfamily: Cytochrome c oxidase subunit II-like, transmembrane region
Family: Cytochrome c oxidase subunit II-like, transmembrane region
Phyre2
| 10 |
|
PDB 2bbj chain B
Region: 373 - 406
Aligned: 34
Modelled: 34
Confidence: 19.5%
Identity: 21%
PDB header:metal transport/membrane protein
Chain: B: PDB Molecule:divalent cation transport-related protein;
PDBTitle: crystal structure of the cora mg2+ transporter
Phyre2
| 11 |
|
PDB 3qnq chain D
Region: 366 - 418
Aligned: 53
Modelled: 53
Confidence: 18.2%
Identity: 9%
PDB header:membrane protein, transport protein
Chain: D: PDB Molecule:pts system, cellobiose-specific iic component;
PDBTitle: crystal structure of the transporter chbc, the iic component from the2 n,n'-diacetylchitobiose-specific phosphotransferase system
Phyre2
| 12 |
|
PDB 3ehb chain B domain 2
Region: 382 - 418
Aligned: 36
Modelled: 37
Confidence: 17.3%
Identity: 14%
Fold: Transmembrane helix hairpin
Superfamily: Cytochrome c oxidase subunit II-like, transmembrane region
Family: Cytochrome c oxidase subunit II-like, transmembrane region
Phyre2
| 13 |
|
PDB 1j4n chain A
Region: 375 - 416
Aligned: 42
Modelled: 42
Confidence: 11.3%
Identity: 10%
Fold: Aquaporin-like
Superfamily: Aquaporin-like
Family: Aquaporin-like
Phyre2
| 14 |
|
PDB 3a0b chain T
Region: 381 - 407
Aligned: 27
Modelled: 27
Confidence: 9.7%
Identity: 19%
PDB header:electron transport
Chain: T: PDB Molecule:photosystem ii reaction center protein t;
PDBTitle: crystal structure of br-substituted photosystem ii complex
Phyre2
| 15 |
|
PDB 3arc chain T
Region: 381 - 407
Aligned: 27
Modelled: 27
Confidence: 9.7%
Identity: 19%
PDB header:electron transport, photosynthesis
Chain: T: PDB Molecule:photosystem ii reaction center protein t;
PDBTitle: crystal structure of oxygen-evolving photosystem ii at 1.9 angstrom2 resolution
Phyre2
| 16 |
|
PDB 2fno chain A domain 1
Region: 49 - 72
Aligned: 24
Modelled: 24
Confidence: 8.7%
Identity: 25%
Fold: GST C-terminal domain-like
Superfamily: GST C-terminal domain-like
Family: Glutathione S-transferase (GST), C-terminal domain
Phyre2
| 17 |
|
PDB 1m57 chain H
Region: 380 - 418
Aligned: 38
Modelled: 39
Confidence: 7.8%
Identity: 16%
PDB header:oxidoreductase
Chain: H: PDB Molecule:cytochrome c oxidase;
PDBTitle: structure of cytochrome c oxidase from rhodobacter2 sphaeroides (eq(i-286) mutant))
Phyre2
| 18 |
|
PDB 1ar1 chain B
Region: 380 - 418
Aligned: 39
Modelled: 39
Confidence: 7.6%
Identity: 10%
PDB header:complex (oxidoreductase/antibody)
Chain: B: PDB Molecule:cytochrome c oxidase;
PDBTitle: structure at 2.7 angstrom resolution of the paracoccus2 denitrificans two-subunit cytochrome c oxidase complexed3 with an antibody fv fragment
Phyre2
| 19 |
|
PDB 1qle chain B
Region: 380 - 418
Aligned: 39
Modelled: 39
Confidence: 7.6%
Identity: 10%
PDB header:oxidoreductase/immune system
Chain: B: PDB Molecule:cytochrome c oxidase polypeptide ii;
PDBTitle: cryo-structure of the paracoccus denitrificans four-subunit2 cytochrome c oxidase in the completely oxidized state3 complexed with an antibody fv fragment
Phyre2
| 20 |
|
PDB 1v54 chain B domain 2
Region: 382 - 417
Aligned: 36
Modelled: 36
Confidence: 6.5%
Identity: 19%
Fold: Transmembrane helix hairpin
Superfamily: Cytochrome c oxidase subunit II-like, transmembrane region
Family: Cytochrome c oxidase subunit II-like, transmembrane region
Phyre2
| 21 |
|
| 22 |
|