Q14524 1295 K
The nsSNP is at position 55 of Pfam domain PF00520.
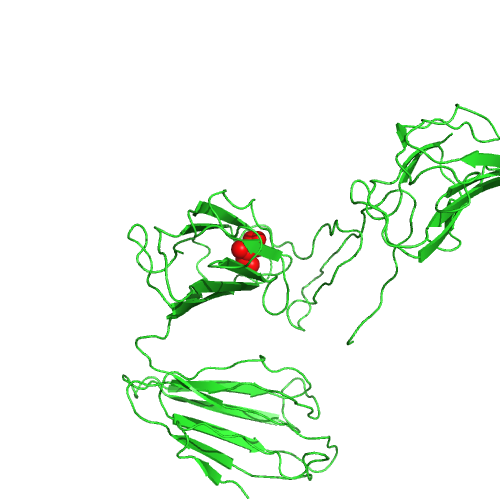
This nsSNP maps to PDB 4dxw, chain D, position 79.
Position predicted to be in a transmembrane helix by TMHMM.
High relative solvent accessibility.
Low conservation (Jensen-Shannon divergence).
K is more favourable than E in the PSSM.
Q14524 has intermediate degree in STRING.
|
Q9H344 134 A
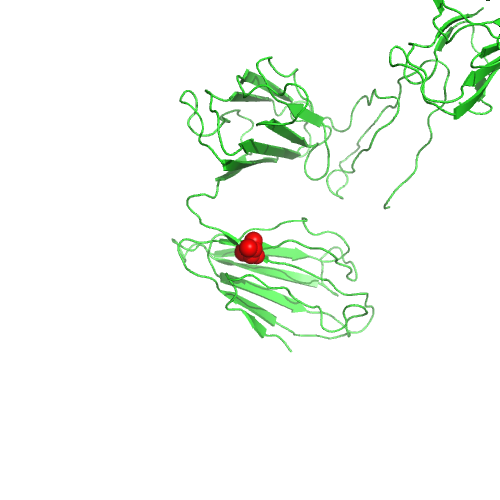
This nsSNP maps to PDB 3uon, chain A, position 133.
Position predicted to be in a transmembrane helix by TMHMM.
High conservation (Jensen-Shannon divergence).
Q9H344 has intermediate degree in STRING.
|
Q9BY79 136 M
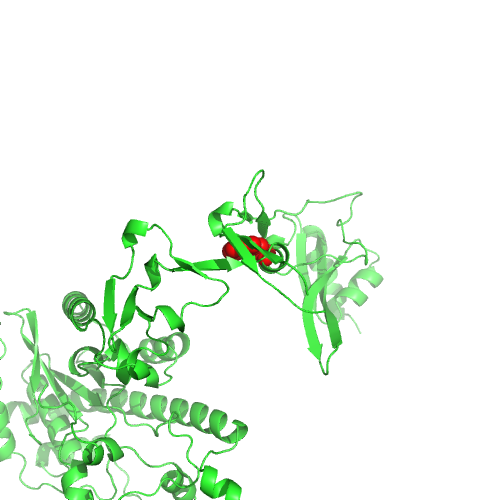
This nsSNP maps to PDB 3kq4, chain D, position 1040.
Low conservation (Jensen-Shannon divergence).
Q9BY79 has low degree in STRING.
|
Q9BY79 182 T
The nsSNP is at position 39 of Pfam domain PF00431.
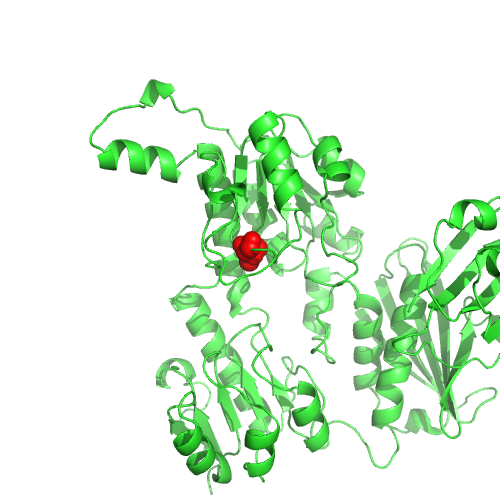
This nsSNP maps to PDB 3kq4, chain D, position 1086.
Low relative solvent accessibility.
T is less favourable than I in the PSSM.
Q9BY79 has low degree in STRING.
|
Q01831 690 S
The nsSNP is at position 7 of Pfam domain PF10404.
This nsSNP maps to PDB 2qsh, chain A, position 496.
Low relative solvent accessibility.
High conservation (Jensen-Shannon divergence).
S is less favourable than W in the PSSM.
Q01831 has intermediate degree in STRING.
|
Q9UK13 519 T
This nsSNP maps to PDB 2csh, chain A, position 23.
Position predicted to be phosphorylated by NetPhos.
High conservation (Jensen-Shannon divergence).
T is less favourable than S in the PSSM.
Q9UK13 has low degree in STRING.
|
Q9UK13 557 R
The nsSNP is at position 10 of Pfam domain PF13465.
This nsSNP maps to PDB 2csh, chain A, position 61.
Loss of Glycine may affect secondary structure.
High conservation (Jensen-Shannon divergence).
R is more favourable than G in the PSSM.
Q9UK13 has low degree in STRING.
|
O43175 261 M
The nsSNP is at position 151 of Pfam domain PF02826.
This nsSNP maps to PDB 1ygy, chain A, position 258.
This position is annotated as NAD binding site.
This position is annotated as putative NAD binding site.
Low relative solvent accessibility.
High conservation (Jensen-Shannon divergence).
M is less favourable than V in the PSSM.
O43175 has high degree in STRING.
|
O43520 454 G
The nsSNP is at position 4 of Pfam domain PF12710.
This nsSNP maps to PDB 2zxe, chain A, position 376.
This position is annotated in UniProt as ACT_SITE.
The nsSNP is at a catalytic site.
High conservation (Jensen-Shannon divergence).
O43520 has intermediate degree in STRING.
|
P98169 760 S
High relative solvent accessibility.
Low conservation (Jensen-Shannon divergence).
S is more favourable than N in the PSSM.
P98169 has low degree in STRING.
|