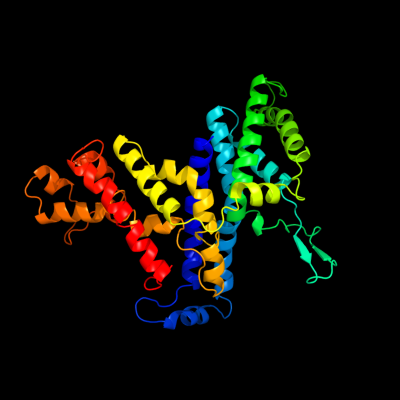
| Secondary structure and disorder prediction | |
 |
| | |
1 | . | . | . | . | . | . | . | . | 10 | . | . | . | . | . | . | . | . | . | 20 | . | . | . | . | . | . | . | . | . | 30 | . | . | . | . | . | . | . | . | . | 40 | . | . | . | . | . | . | . | . | . | 50 | . | . | . | . | . | . | . | . | . | 60 |
| Sequence | |
M | L | S | Q | I | Q | R | F | G | G | A | M | F | T | P | V | L | L | F | P | F | A | G | I | V | V | G | L | A | I | L | L | Q | N | P | M | F | V | G | E | S | L | T | D | P | N | S | L | F | A | Q | I | V | H | I | I | E | E | G | G |
| Secondary structure | |
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|  |  |  |  |  |  |  |
|
|
|
|
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |
| SS confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder | |
? | ? | ? |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| ? | ? |
| ? | ? | ? | ? | ? | ? | ? |
| ? | ? | ? |
| ? |
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
| | |
. | . | . | . | . | . | . | . | . | 70 | . | . | . | . | . | . | . | . | . | 80 | . | . | . | . | . | . | . | . | . | 90 | . | . | . | . | . | . | . | . | . | 100 | . | . | . | . | . | . | . | . | . | 110 | . | . | . | . | . | . | . | . | . | 120 |
| Sequence | |
W | T | V | F | R | N | M | P | L | I | F | A | V | G | L | P | I | G | L | A | K | Q | A | Q | G | R | A | C | L | A | V | M | V | S | F | L | T | W | N | Y | F | I | N | A | M | G | M | T | W | G | S | Y | F | G | V | D | F | T | Q | D |
| Secondary structure | |
 |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
|
|
|
|
|
|
|
|  |  |
| SS confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| ? | ? |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| ? | ? | ? | ? |
| ? | ? | ? | ? |
| ? | ? | ? | ? | ? |
| Disorder confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
| | |
. | . | . | . | . | . | . | . | . | 130 | . | . | . | . | . | . | . | . | . | 140 | . | . | . | . | . | . | . | . | . | 150 | . | . | . | . | . | . | . | . | . | 160 | . | . | . | . | . | . | . | . | . | 170 | . | . | . | . | . | . | . | . | . | 180 |
| Sequence | |
A | V | A | G | S | G | L | T | M | M | A | G | I | K | T | L | D | T | S | I | I | G | A | I | I | I | S | G | I | V | T | A | L | H | N | R | L | F | D | K | K | L | P | V | F | L | G | I | F | Q | G | T | S | Y | V | V | I | I | A | F |
| Secondary structure | |
 |
|
|
|
|
|
|  |  |  |  |
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
|
|  |  |  |  |  |  |
|
|
|
|  |  |  |  |  |  |  |
| SS confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder | |
| ? |
|
|
|
|
|
|
|
|
|
|
|
|
| ? | ? |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
| | |
. | . | . | . | . | . | . | . | . | 190 | . | . | . | . | . | . | . | . | . | 200 | . | . | . | . | . | . | . | . | . | 210 | . | . | . | . | . | . | . | . | . | 220 | . | . | . | . | . | . | . | . | . | 230 | . | . | . | . | . | . | . | . | . | 240 |
| Sequence | |
L | V | M | I | P | C | A | W | L | T | L | L | G | W | P | K | V | Q | M | G | I | E | S | L | Q | A | F | L | R | S | A | G | A | L | G | V | W | V | Y | T | F | L | E | R | I | L | I | P | T | G | L | H | H | F | I | Y | G | Q | F | I |
| Secondary structure | |
 |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
|
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|  |  |  |  |  |  |  |  |  |
| SS confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
| | |
. | . | . | . | . | . | . | . | . | 250 | . | . | . | . | . | . | . | . | . | 260 | . | . | . | . | . | . | . | . | . | 270 | . | . | . | . | . | . | . | . | . | 280 | . | . | . | . | . | . | . | . | . | 290 | . | . | . | . | . | . | . | . | . | 300 |
| Sequence | |
F | G | P | A | A | V | E | G | G | I | Q | M | Y | W | A | Q | H | L | Q | E | F | S | L | S | A | E | P | L | K | S | L | F | P | E | G | G | F | A | L | H | G | N | S | K | I | F | G | A | V | G | I | S | L | A | M | Y | F | T | A | A |
| Secondary structure | |
 |  |  |  |  |  |  |
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
| SS confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| ? | ? | ? |
|
|
|
| ? | ? |
|
| ? |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
| | |
. | . | . | . | . | . | . | . | . | 310 | . | . | . | . | . | . | . | . | . | 320 | . | . | . | . | . | . | . | . | . | 330 | . | . | . | . | . | . | . | . | . | 340 | . | . | . | . | . | . | . | . | . | 350 | . | . | . | . | . | . | . | . | . | 360 |
| Sequence | |
P | E | N | R | V | K | V | A | G | L | L | I | P | A | T | L | T | A | M | L | V | G | I | T | E | P | L | E | F | T | F | L | F | I | S | P | L | L | F | A | V | H | A | V | L | A | A | S | M | S | T | V | M | Y | L | F | G | V | V | G |
| Secondary structure | |
 |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|
|
|  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |  |
|  |  |  |
| SS confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Disorder confidence | |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
| | |
. | . | . | . | . | . | . | . |
| Sequence | |
N | M | G | G | G | L | I | D |
| Secondary structure | |
|
|
|
|
|
|
|
|
| SS confidence | |
|
|
|
|
|
|
|
|
| Disorder | |
|
|
|
| ? | ? | ? | ? |
| Disorder confidence | |
|
|
|
|
|
|
|
|
| |
| Confidence Key |
| High(9) | |
|
|
|
|
|
|
|
|
|
Low (0) |
| ? | Disordered |
  | Alpha helix |
  | Beta strand |
Hover over an aligned region to see model and summary info
Please note, only up to the top 20 hits are modelled to reduce computer load
|
| 1 |
|
PDB 3qnq chain D
Region: 3 - 365
Aligned: 347
Modelled: 359
Confidence: 100.0%
Identity: 15%
PDB header:membrane protein, transport protein
Chain: D: PDB Molecule:pts system, cellobiose-specific iic component;
PDBTitle: crystal structure of the transporter chbc, the iic component from the2 n,n'-diacetylchitobiose-specific phosphotransferase system
Phyre2
| 2 |
|
PDB 1hgv chain A
Region: 41 - 83
Aligned: 43
Modelled: 43
Confidence: 62.2%
Identity: 14%
PDB header:virus
Chain: A: PDB Molecule:ph75 inovirus major coat protein;
PDBTitle: filamentous bacteriophage ph75
Phyre2
| 3 |
|
PDB 1j9i chain A
Region: 303 - 328
Aligned: 16
Modelled: 26
Confidence: 16.1%
Identity: 25%
Fold: Putative DNA-binding domain
Superfamily: Putative DNA-binding domain
Family: Terminase gpNU1 subunit domain
Phyre2
|
| Detailed template information | |
Due to computational demand, binding site predictions are not run for batch jobs
If you want to predict binding sites, please manually submit your model of choice to 3DLigandSite
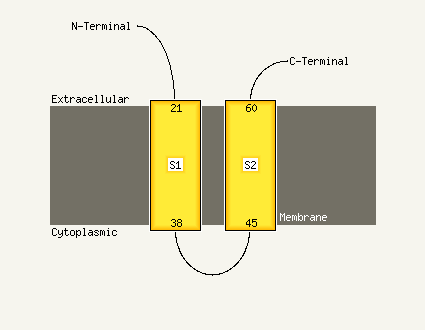
| Transmembrane helix prediction | |
Transmembrane helices have been predicted in your sequence to adopt the topology shown below
Phyre is for academic use only
| Please cite: Protein structure prediction on
the web: a case study using the Phyre server |
| Kelley LA and Sternberg MJE. Nature Protocols
4, 363 - 371 (2009) [pdf] [Import into BibTeX] |
| |
| If you use the binding site
predictions from 3DLigandSite, please also cite: |
| 3DLigandSite: predicting ligand-binding sites using similar structures. |
| Wass MN, Kelley LA and Sternberg
MJ Nucleic Acids Research 38, W469-73 (2010) [PubMed] |
| |
|
|
|
|